Imaging/Flow
Flow Cytometry (FCRG)
Mission
The Flow Cytometry Research Group was formed in 2012 by Peter Lopez (NYU) and Scott Tighe (UVM) with the goal of providing information related to the art of flow cytometry, including its cross-technology applications to genomics, proteomics, and other core-related areas.
- Mehrnoosh Abshari, NIH/NIDCR
- Roxann Ashworth (Executive Board liaison) - Johns Hopkins University, School of Medicine
- Claudia Bispo, University of California San Francisco
- Sara Bowen, Dignity Health St. Joseph's Hospital and Medical Center
- Ching-Yuan (Steve) Chen, Columbia University Irving Medical Center
- Xiaoxuan Fan, University of Maryland School of Medicine
- Kevin Ferro, Stowers Institute for Medical Research
- Christiane Hassel, Indiana University
- Celine Lages, Cincinnati Children's Hospital Medical Center
- Pam Moody, Cold Spring Harbor Laboratory
- Steven Polter, University of Rochester Medical Center
- Kenneth Quayle, Cincinnati Children's Hospital Medical Center
- Kathy Schaefer, HHMI Janelia Research Campus
- Rachael Sheridan (Co-Chair) - Van Andel Research Institute Flow Cytometry Core
- Jane Srivastava (Chair) - Gladstone Institutes
- John Tigges, Beth Israel Deaconess Medical Center
- Eric Wieder, University of Miami Miller School of Medicine
The Flow Cytometry Research Group (FCRG) focuses on flow cytometry-related projects. In many experimental workflows, cells sorted in a flow cytometry core are further analyzed in other cores. Currently, the RG emphasizes exploration of the various effects that cell sorting may or may not have on sorted material and thus the effects on downstream analysis by other shared facilities.
CURRENT STUDY
DROP DELAY
It is known that correct drop delay calibration and accurate sort recovery is always critical when sorting. The FCRG's current study will test drop delay settings in sorters with options to set drop delay manually and those with fully automated, non-adjustable drop delay set-up. Data collected will help to determine the accuracy of drop delay and sort recovery and purity for both types of sorters, and help serve as a guide for sort operators to assist in determining drop delay accuracy.
Poster presented at 2023 CYTO and ABRF Annual Meetings:
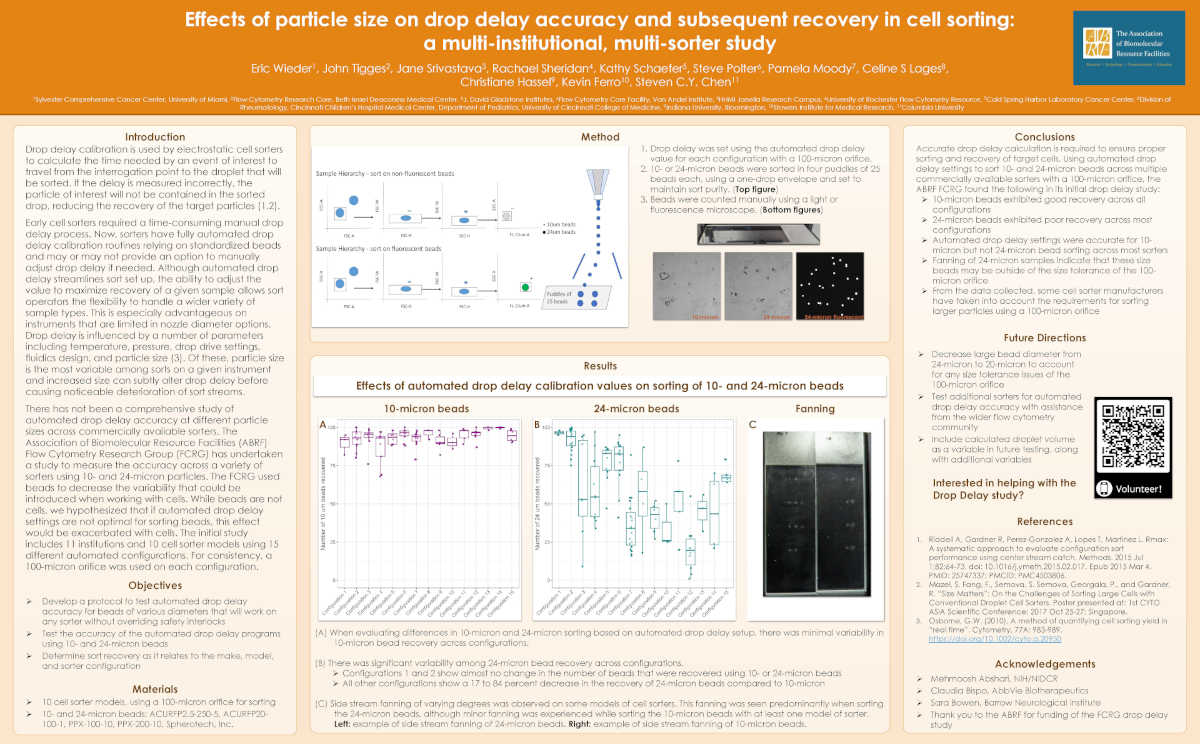
PAST STUDIES/PROJECTS
FCRG Community Survey (2021)
GENE EXPRESSION AFTER SORTING
The first study proposed by the group investigated the impact of cell sorting on gene expression. Variables such as sorting versus no sorting, high versus low pressure, and the presence versus absence of UV light were analyzed with RNA-seq and microarray in Jurkat cells, primary B cells, and mouse ES cells. A manuscript with this study’s results is being prepared.
SORTER CLEANLINESS
Cytometry facilities around the world were surveyed regarding their sorter cleaning practices. A subset of facilities submitted samples for testing. Most sorters had significant endotoxin contamination, but little to no RNase. Bacterial concentrations ranged from none to substantial. There was no correlation found between sorter cleanliness and any surveyed facility variables, including sorter age, cleaning practices, date of last PM, sheath source, or known recent contamination. The results will be published in combination with the endotoxin removal results.
ENDOTOXIN REMOVAL
Since a number of sorters assayed in the sorter cleanliness study were contaminated with endotoxin, effectiveness of an H2O2 cleaning procedure was tested to assess removal of the endotoxin from the sorters. Sheath samples collected before and after cleaning were tested with a LAL quantitation kit and it was determined that the contamination was only partially mitigated. Also, the endotoxin levels reached pre-cleaning levels within a few weeks of the sterilization. The results will be published alongside the sorter cleanliness survey results.
- ABRF 2022 workshop - Caveats and Best Practices for Study Design and Data Analysis in High-dimensional Flow
- ABRF 2019 poster - A Multi-Core Study on How Different Fixation Methods Prior to Sorting Impact the Purity, Quality, and Yield of RNA From Sorted Cells
- ABRF 2018 Scientific Session - Single Cell Sorting and the Bioinformatics Pathway
- ABRF 2018 Satellite Workshop - Bridging the Gap: Isolation to Translation (Single-Cell RNA-Seq)
- Basics of RNA-Seq by Michael Kelly
- Methods, Applications & Analysis of scRNA-Seq: How an Integrated Understanding of Every Step Makes for a Better Single Cell Experiment by Michael Kelly
- Open Workflows to Enable and Understand Single-Cell RNA-Seq Data by Nathan Salomonis
- ABRF 2017 poster - Endotoxin Contamination of Cell Sorters: Evaluating Cleaning and Testing Procedures
- ABRF 2017 - Sheath Contamination Survey: An Examination of Common Laboratory Practices
Presentation by Dave Adams
Poster - ABRF 2016 - Evaluating the Effects of Cell Sorting on Gene Expression
Presentation by Andrew Box
Poster - ABRF 2016 poster - Evaluating Cell Sorter Cleaning Procedures Across ABRF-FCRG Institutions by Testing for Common Contaminants
- ABRF 2015 - Evaluating the Effects of Cell Sorting on Gene Expression
Presentation
Poster - ABRF 2014 poster - Evaluating Effects of Cell Sorting on Cellular Integrity and Gene Expression
- ABRF 2013 presentation by Scott Tighe - Overview of the New FCRG and Proposed Cell Sorting (FACS) Microarray Study
- ABRF 2013 poster - The New ABRF Flow Cytometry Research Group
| 1) | This paper has a detailed protocol for how to properly prepare a sorter for RNA isolation and sort cells with Trizol, RLT, and RNase inhibitor. It also includes an estimated yield of RNA from certain cell types. - View Document (802K) |
| Member Name | Organization | Details |
|---|---|---|
| Andrew C. Box | Stowers Institute for Medical Research | Member: 11/12 - 03/16 Co-chair: 05/15 - 03/16 Ad hoc: 04/16 - 04/21 |
| Dr Sridar V Chittur | SUNY Albany | Member: 11/12 - 05/15 |
| Monica DeLay | CCHMC | Member: 09/12 - 03/17 Co-chair: 03/13 - 03/16 Ad-hoc: 04/17 - 04/21 |
| Maris Handley | Massachusetts General Hospital | Member: 04/15 - 03/17 |
| Peter Lopez | NYU Office Collaborative Science | Member: 09/12 - 03/15 Ad hoc: 04/15 - 04/21 |
| Dr Thomas Neubert | New York Univ Sch of Med | EB Liaison: 09/12 - 03/13 |
| Mr. Hank Pletcher | University of Pennsylvania | Ad hoc: 11/12 - 03/13 |
| Scott Tighe | Vermont Cancer Center | Chair- RG Co-Founder: 02/12 - 03/13 Member, ad hoc-co-chair: 04/13 - 05/15 |
| Paula B Turpen | University of Nebraska Medical Center | EB Liaison: 04/13 - 04/14 |
| Frances Weis-Garcia | Memorial Sloan Kettering Cancer Center | EB Liaison: 05/14 - 03/17 |
Questions or interest in joining an ABRF research group? Contact us!
Light Microscopy (LMRG)
Mission
Our goal is to promote scientific exchange between researchers, specifically those in core facilities in order to increase our general knowledge and experience. We seek to provide a forum for multi-site experiments exploring “standards” for the field of light microscopy.
- Benjamin Abrams - University of California Santa Cruz
- Constadina Arvanitis - Northwestern University
- Linda Callahan - University of Rochester Medical Center
- Richard Cole - Wadsworth Center
- Natalia Dworak - University of Virginia
- Joseph Dragavon - University of Colorado
- Corinne Esquibel - Van Andel Institute
- Kari Herrington - University of California, San Francisco
- Michelle Itano - University of North Carolina
- Justine Kigenyi (EB liaison) - Kansas University Medical Center
- Soyeon Kim - University of California, San Francisco
- Kristopher Kubow - James Madison University
- DeLaine Larsen - University of California, San Francisco
- Guillermo Marques - University of Minnesota
- Arvydas Matiukas - SUNY Upstate Medical University
- Valeria Mezzano, NYU School of Medicine
- Thomas Pengo - University of Minnesota
- Josh Rappoport - Boston College
- Mark Sanders - University of Minnesota
- Jian Wei Tay - University of Colorado
- Erika Wee - Cold Spring Harbor Laboratory
3D Image Analysis Tools and Reproducibility Event
Wednesday, April 27, 2022
Workshop breakout session recordings:
Nearly 100 attendees participated in this session on 3D image analysis tools and how they can help improve reproducibility, hosted by the Light Microscopy Research Group.
This event was centered around the LMRG’s ongoing study of reproducibility in 3D image analysis in which we seek to characterize and identify sources of (ir)reproducibility in 3D segmentation. Representatives from several major 3D analysis platforms walked attendees through how to segment and analyze the images from our study and discussed how their platforms support reproducible image analysis.
This program was targeted to people who are interested in learning more about a specific image analysis platform, learning how to use an image analysis tool better, and/or learning about how reproducibility is addressed by these platforms.
If you have questions about the LMRG study, please contact LMRG member Jessica Hornick for more information.
New LMRG Study - Volunteers Needed!
The Light Microscopy Research Group of the ABRF is launching a study to assess reproducibility in quantitative image analysis and needs YOUR help! We are seeking volunteers to segment 3D fluorescence microscope image sets and provide us with both their analysis and their analysis strategy. Our goal is not to find novel segmentation algorithms, but to compare segmentation results and strategies across a broad cross-section of volunteer analysts.
You can help by analyzing one (or more!) images for us! We need participants from all levels of image analysis experience. Everyone can participate - from students to core directors to image analysis experts. See our study website to find-out more: https://sites.google.com/view/lmrg-image-analysis-study
November 2020 ABRF LMRG/Industry Partner Discussion
The Light Microscopy Research Group and Corporate Relations Committee organized a conversation between light microscopy user groups and corporate partners, including Leica, Nikon, Olympus and Zeiss. This conversation provided an opportunity for each company to explain how they are being impacted by the COVID-19 pandemic and how they are adapting to the evolving needs of their customers.
Date: 11/18/2020
Time: 1:00-2:30 EST
View session recording.
Participants:
Ben Abrams, ABRF Light Microscopy Research Group and Corporate Relations Committee member, Director - Life Science Microscopy Facility Univ. of California Santa Cruz
Rich Cole, ABRF 2020 President, Light Microscopy Research Group member, Director - Advanced Light Microscopy & Image Analysis Core, New York State Dept. of Health Wadsworth Center
Joshua Rappoport, ABRF Light Microscopy Research Group member, Executive Director - Research Infrastructure at Boston College, Secretary – Core Technologies for Life Sciences
Michelle Itano, Director, UNC Neuroscience Microscopy Core Facility
Leica:
Greg Eppink General Manager Microscopy
Ryan Hrejsa, Senior Marketing Manager
Nikon:
Mike Gallo, General Manager, Service
Mike Johnson, Senior Regional Manager, Sales
Lynne Chang, Senior Marketing Manager
Olympus:
Shane Andrews, Manager, Sales Research Market, Life Science, Olympus Corporation of the Americas Scientific Solutions Group
Kerry Israel, Manager, Marketing Communications, Life Science, Olympus Corporation of the Americas Scientific Solutions Group
Zeiss:
Joseph Huff, Head of Marketing, North America, Zeiss Research Microscopy Solutions
Rosey Manser, Head of Business Development for Core Facilities
Studies
Activities
| 1) | ABRF2014 LMRG Group and CCMA Travel Award Winners - Richard Cole, John Russ, and Claire Brown (2,464K) - CCMA Travel Award Winners (1,934K) |
| 2) | ABRF2013 LMRG Photo - ABRF2013 LMRG Photo |
| 3) | ABRF 12 RG group photo - 2012 Meeting Group Photo |
| 4) | ABRF 12 RG intro talk - LMRG 2012 Overview Talk |
| 5) | ABRF 12 RG data talk - Study #2 Presentation (7,190K) |
| 6) | Proteomics workshop-- Get on Your Way to Microproteomics with laser Microdissection - Proteomics - Sarah Baxter |
| 7) | Proteomics workshop-- Sample prep - Tissue Proteomics Sample Preparation Leica Talk |
| 8) | ABRF 11 LMRG group photo - View Document (3,315K) |
| 9) | ABRF 11 RG talk - View Document (13,284K) |
| 10) | ABRF 2010 talk - View Document (4,643K) |
| 11) | Talk for ABRF 09 - View Document (10,567K) |
Protocols
| 1) | This is a paper written by Cole and Brown and gives a lot of background on how to measure point spread functions, why you may want to measure them, how to prepare bead slides and then a detailed protocol for setting up the instrument and measuring and interpreting the PSF. This article should be looked at first before referring to the specific protocols for the individual microscope platforms. - Nature Protocols Paper (1,371K) |
| 2) | PSF Protocol Leica SP5 - Measuring the PSF with the 0.175 um green fluorescent bead sample. - PSF Protocol Leica SP5 |
| 3) | PSF Protocol Nikon A1 - Measuring the PSF with the 0.175 um green fluorescent bead sample. - PSF Protocol Nikon A1 |
| 4) | PSF Protocol Olympus FV1000 - Measuring the PSF with the 0.175 um green fluorescent bead sample. - PSF Protocol Olympus FV1000 |
| 5) | PSF Protocol Zeiss 510 - Measuring the PSF with the 0.175 um green fluorescent bead sample using the AIM software. - PSF Protocol Zeiss 510 |
| 6) | Spectral Accuracy Protocol Leica SP5 - A mirror slide will be provided and you will look at the reflections of the laser lines into the spectral detector to determine if the wavelength readouts are accurate. - Spectral Accuracy Protocol Leica SP5 |
| 7) | Spectral Accuracy Protocol Olympus FV1000 - A mirror slide will be provided and you will look at the reflections of the laser lines into the spectral detector to determine if the wavelength readouts are accurate. - Spectral Accuracy Protocol Olympus FV1000 |
| 8) | Spectral Accuracy Protocol Zeiss 710 - A mirror slide will be provided and you will look at the reflections of the laser lines into the spectral detector to determine if the wavelength readouts are accurate. Uses the ZEN software. - Spectral Accuracy Protocol Zeiss 710 (261K) |
| 9) | Spectral Accuracy Protocol Zeiss 510 - A mirror slide will be provided and you will look at the reflections of the laser lines into the spectral detector to determine if the wavelength readouts are accurate. Uses the AIM software. - Spectral Accuracy Protocol Zeiss 510 |
| 10) | Spectral Separation Accuracy Protocol Leica SP5 - Protocol for spectral separation testing of software un-mixing with double orange stained beads. - Spectral Separation Accuracy Protocol Leica SP5 - Bead Core Spectra Text File - Bead Ring Spectra Text File - Bead Core Spectra lsf File (12K) - Bead Ring Spectra lsf File (9K) |
| 11) | Protocol for testing the spectral separation accuracy of the Olympus FV1000. - Spectral Separation Protocol Olympus FV1000 (1,133K) |
| 12) | Study #1 Laser Stability, Alignment and Co-registration Protocol - Study #1 Laser Stability, Alignment and Co-registr |
| 13) | Improved GFP imaging in live cells - Improved GFP imaging in live cells |
Publications
-
Microsc and Microanal. 2013 Dec;19(6):1653-68.Cole, RW, Thibault, M, Bayles, CJ, Eason, B, Girard, AM, Jinadasa, T, Opansky, C, Schulz, K, Brown, CM,
-
Microsc Microanal. 2011 Aug;17(4):598-606.Stack RF, Bayles CJ, Girard AM, Martin K, Opansky C, Schulz K, Cole RW.Microsc Microanal.
-
Nature Protocols 6 (2011):1929RW Cole, T Jinadasa, CM Brown
| Member Name | Organization | Details |
|---|---|---|
| Pamela Scott Adams | Trudeau Institute | Ad hocEB Liaison: 02/09 - 03/10 |
| Carol J Bayles | Cornell University | Member: 04/08 - 05/15 |
| Claire M Brown | McGill University | Chair: 03/11 - 04/15 Member: 09/10 - 03/18 |
| Lisa Cameron | Duke University | Member |
| Richard Cole | Wadsworth Center | Chair: 03/08 - 03/11 |
| Arnold M. Falick | HHMI-UC Berkeley | Ad hocEB Liaison: 03/08 - 02/09 |
| Paul Furcinitti | UMass Medical School | Member: 05/13 - 12/14 |
| Anne-Marie Girard | Center for Genome Research and Biocomputing, Oregon State University | Member: 02/10 - 05/15 |
| William G Hendrickson | Univ. of Illinois, Chicago | EB Liaison: 09/13 - 04/14 |
| Jessica Hornick | Northwestern University | Member through 12/22 |
| Karen R Jonscher | University of Colorado Denver | Ad hocEB: 04/10 - 03/13 |
| Gary Laevsky | Princeton University | Member: 01/17-04/19 |
| Karen Martin | West Virginia University | Member: 12/08 - 05/15 |
| George McNamara | U Miami | Member: 06/11 - 03/13 Ad hoc: 03/13 - 12/14 |
| Kary Oakleaf | Molecular Probes, Life Technologies | Member: 06/13 - 12/14 |
| Cynthia Opansky | Blood Center of Wisconsin | Member: 02/09 - 12/13 |
| James Powers | Indiana University | Member through 12/22 |
| Joshua Rappoport | Northwestern University | Member: 03/15-01/19 |
| Katherine Schulz | Blood Center of Wisconsin | Member: 03/09 - 04/13 |
| Robert F. Stack | Wadsworth Center NYSDOH | Member: 02/09 - 03/11 |
| Marc Thibault | Ecole Polytechnique | Member: 06/11 - 03/14 |
| Erika Wee | McGill University | Chair: 04/15 - 05/18 |
| Frances Weis-Garcia | Memorial Sloan Kettering Cancer Center | EB Liaison: 05/14 - 04/15 |
Questions or interest in joining an ABRF research group? Contact us!
